A Factored Co-morbidity Approach for Modelling CKD Progression
An absract published in the British Renal Society 2017 (BRS2017)
Introduction
One of the key challenges in understanding CKD progression is its multifaceted aetiology. This is evident as it is commonly observed that hypertension, heart disease and diabetes are common co-morbidities of CKD. In other words, the existence of co-morbidities can potentially alter the risk of CKD progression, e.g. from stage 3 to 5. Unfortunately, “flat” risk models such as logistic regression, e.g. as implemented by the QKidney score and many similar risk models, are not designed to extract the rich structure induced by a multitude of co-morbidities, the state of which are often captured in routinely collected patient data.
Methods
We revisit the modelling of CKD progression by proposing a Bayesian network that has a certain regular structure, which has attractive properties such as tractability and efficient computation. The risk of progression to more severe CKD stages can be inferred not only from eGFR but signs and symptoms (s) related to kidney impairment, while causality of CKD is captured by risk factors () and treatment strategies (). Each co-morbidity is then allowed to directly interact with the CKD progression risk only if the diagnosis of the co-morbidity is available. Where this information is not available, the state of the co-morbidity can be inferred from its own and variables.
In addition, the model can also incorporate past episodes of acute kidney injury (AKI) when estimating the risk of CKD progression. This exploits the fact that people with AKI are more likely to have a lower eGFR value. The extent of this degradation is explicitly modelled by the Bayesian network. Similar to the above, as before, when the AKI information is not available, the risk of AKI can be inferred from its own and variables, in a process known as ‘marginalisation’. Therefore, the interplay between CKD and AKI can be captured by the model.
Analysis
As a proof of concept, the proposed Bayesian network can be verified “by parts”. In essence, the factored co-morbid approach offers a divide-and-conquer strategy to implement risk models. Some of the relationships of the model are informed by the literature and can be expressed in the conditional probabilities below.
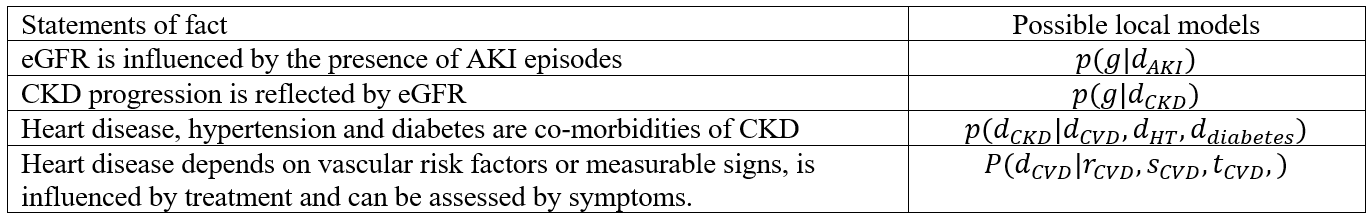

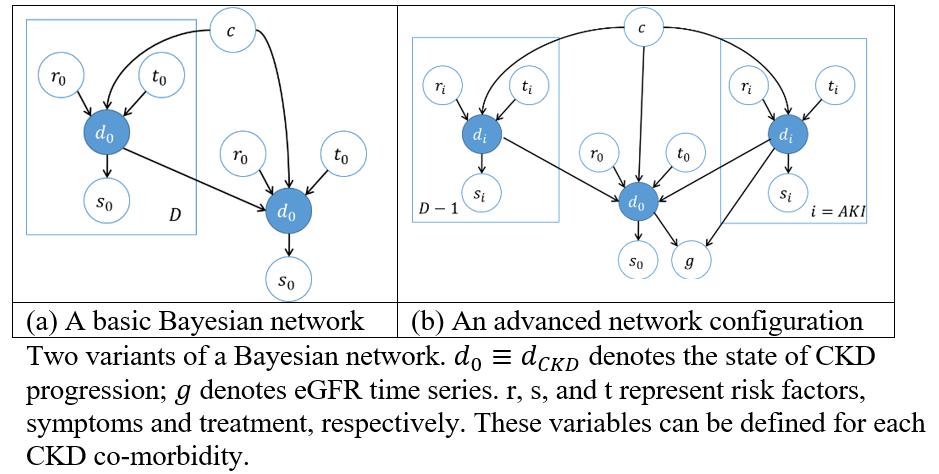
The local models can be linked to form a single Bayesian network as shown in figure (a) below. In (b), the model is tailored for AKI, which makes provision for eGFR () to be influenced by AKI directly.
Conclusion and relevance
Current risk models are ‘flat’ and do not exploit the rich information captured in patient records. The proposed factored co-morbidity approach offers a natural next step whilst being tractable and efficient in computation. The merit of this model will be investigated as part of the MRC Modelling CKD project using data extracted from the Royal College of GPs database.
What’s next?
Download the abstract here and the poster here.
Cite this blog post
Bibtex
@misc{ poh_2018_01_13_BRS2017-bayesian-justification,
author = {Norman Poh},
title = { A Factored Co-morbidity Approach for Modelling CKD Progression },
howpublished = {\url{ http://normanpoh.github.com/blog/2018/01/13/BRS2017-bayesian-justification.html},
note = "Accessed: ___TODAY___"
}